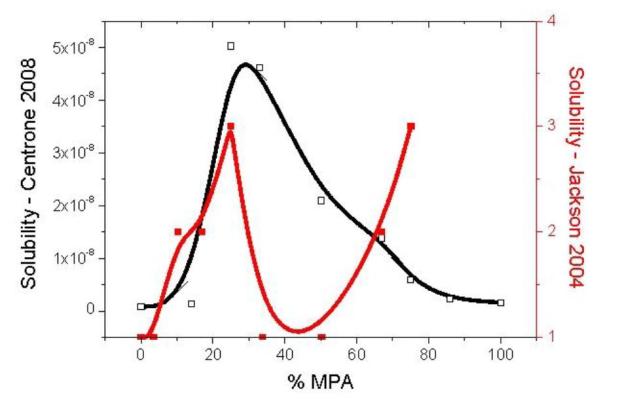
Well nearly. Possibly as close as we might get.
If you have followed the Spherical Nucleic Acid / SmartFlare / StickyFlare story on this blog, you will know that we have raised doubts about the endosomal escape of these nanoparticles which are supposed to reach the cytosol of cells where they could detect mRNAs. We have even published an article on this topic. The Mirkin group has developed the technology and it has been commercialized four years ago for the live cell detection of mRNAs by EMD Millipore.
One PNAS paper on the topic was Briley et al (see here for letter to PNAS and what happened to that letter). In his PhD dissertation (Briley, W. E. (2016), William Edward Briley respond to our criticism. I reproduce the relevant section below. One might note that there is an incredibly simple way to determine the localisation of these particles inside the cells: electron microscopy, a technique which has been used for this exact purpose for over 50 years. We have used it. The results were unambiguous.
2.3 Commentary on the Endosomal Escape of SNAs
Though the endosomal escape of SNA nanostructures such as the Nanoflare and stickyflare is evident based upon their ability to provide sequence-specific information regarding RNA levels and locations within cells, one researcher has concluded that SNAs cannot escape from endosomes.[75] That researcher [That’s me!] is ignoring the many papers now that use such architectures for sequence-specific cell-sorting experiments. Indeed, if these architectures, which are taken up by scavenger-receptor mediated endosomal pathways,[68a] do not escape the endosome, then it is difficult to understand the reports by the many groups who have documented the sequencespecific function of SNAs (all of which require endosomal escape), in antisense gene regulation,[12, 44-45, 53] siRNA gene regulation,[20, 68b, 76] nanoflare gene detection,[47, 67a, 67c, 77] and sticky-flare gene detection.[78] Perhaps the best demonstration of this sequence specificity is in the function of the multiplexed Nanoflare.[67a] This nanoconjugate, discussed in chapter 1, contains two targeting sequences specific to two different genes (actin and survivin). When cells treated with multiplexed nanoflares were subjected to siRNA that specifically knocked down the expression of survivin, a corresponding decrease of fluorescence was observed in the nanoflare’s survivin-associated fluorescence (cy5), but not the actin-associated fluorescence (cy3) compared to control.[67a] Likewise, when cells were subjected to actin-targeting siRNA, a decrease in actin-associated fluorescence was observed, with no decrease of survivin-associated fluorescence.
These results indicate that the detection by nanoflares is unique to the targeted mRNA transcript. To rule out any possible effects of the fluorophores, the Cy5 and Cy3 dyes were switched to the other gene (actin-cy5, survivin-cy3), and again the corresponding sequence-specific responses were observed. Importantly, since the development of the multiplexed nanoflare, other research groups have independently developed nanoflare-like structures capable of sequence-specific detection of 3,[57] and even 4[79] genes simultaneously. Additionally, since the commercialization and sale of the nanoflare platform under the trade name Smartflare (Millipore), dozens of researchers around the world have participated in successful sequence-specific gene detection.[80]
Further evidence of SNA endosome escape can be seen visually through analysis of the sticky-flare construct. If sticky-flares were incapable of escaping endosomes, one would expect consistent colocalization with endosomes. However, this is not the case. The patterns of β-actin targeted sticky-flares, when used in HeLa cells, instead exhibit localization around mitochondria. If such structures were limited to endosomes, it is inconceivable that they would co-localize with specific organelles. This is consistent with mitochondrial localization of multiple RNAs which
has also been identified by other groups, using multiple techniques, in HeLa cells.[74] Sticky-flare release from the endosome was further confirmed by designing sticky-flares targeting a second sequence, a U1 short nuclear RNA (snRNA). U1 snRNA is known to traffic from the cytoplasm to the nucleus. Indeed, cells treated with U1-targeting sticky-flares exhibited specific fluorescence within the nucleus. The pattern observed indicates nuclear localization through endosomal escape and sequence-specific tagging of an RNA that is actively transported into the nucleus. Again, this would not be possible if such structures were confined exclusively to endosomes. Speculation that SNAs do not escape endosomes has been fueled by the observation that the fluorescence pattern of β-actin in MEFs is punctate, which has been interpreted as an indication that the sticky-flares are simply trapped in endosomes. However, β-actin is well known to exhibit punctate fluorescence in many cases, an observation made by others in multiple cell lines, including MEFs.[81] Punctate fluorescence is very common in RNA-labeling studies and well known by researchers familiar with FISH.82 This is due to the fact that RNA is often packaged into large RNA-containing granules, which facilitates transport and translational control of the included transcripts.[82b, 83] Such packaging into granules has been extensively studied using β-actin mRNA. Thus, the fact that sticky-flares targeting β-actin were packaged into granules as previously observed, while U1-targeting sticky-flares were specific to the nucleus in the same cell line, demonstrates the functionality of the construct.
Taken together, the success of the many groups who use flare architectures for the detection and knockdown of RNA in cells, and the work of dozens of labs studying related nanoparticle constructs, provide unambiguous evidence of the ability for such architectures to escape the endosome and participate in reactions exclusive to the cytosol. The mechanism of endosomal escape for nanoparticle-based vehicles is currently unknown and is an interesting and important question that is actively being investigated by many in the field.[84]


Briley, W. E. (2016). Investigation and manipulation of the local microenvironment of spherical nucleic acid nanoconjugates (Order No. 10117274). Available from ProQuest Dissertations & Theses Global. (1795522748). Retrieved from https://search.proquest.com/docview/1795522748?accountid=12117